Community-based antibiotic screening method for polymicrobial infections (2020-090)
An improved antibiotic susceptibility test (AST) that determines minimal inhibitory concentration (MIC) with high physiological relevance for polymicrobial infections.
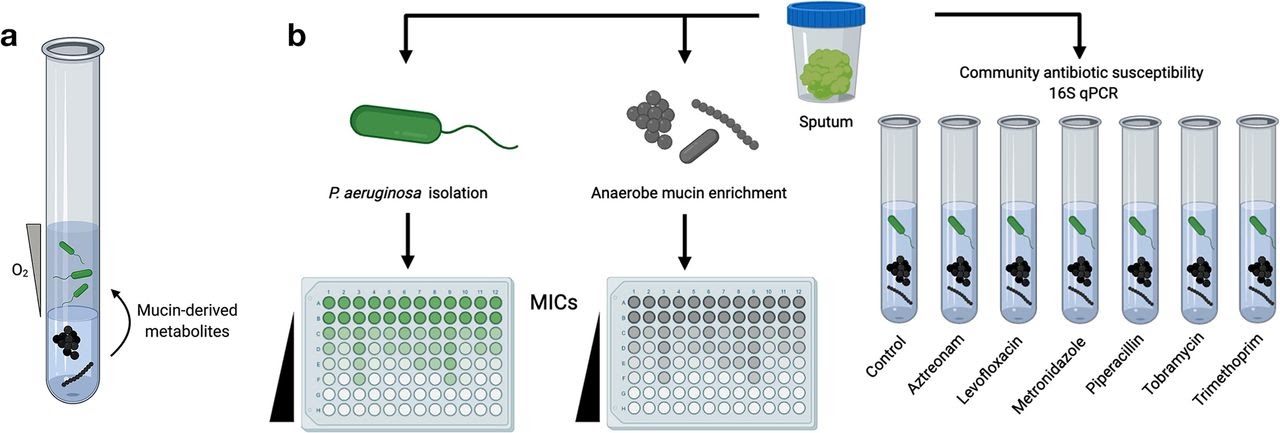
Applications
- Determine physiologically relevant antibiotic minimum inhibitory concentrations (MICs) for polymicrobial infections present in diseases such as cystic fibrosis (CF), rhinosinusitis, chronic obstructive pulmonary disorder (COPD), chronic wounds, and many more.
- High-throughput screening for novel antibiotic treatments for polymicrobial infections.
Key Benefits & Differentiators
- Increased physiological relevance: Accurately models in vivo conditions in patients by utilizing polymicrobial growth conditions.
- Improved antibiotic accuracy and efficacy: Screen and generate personalized, efficacious antibiotic regimens derived from actual patient samples.
- Applicable to a variety of polymicrobial infections: Can be applied to a variety of diseases with polymicrobial infection such as cystic fibrosis (CF), rhinosinusitis, chronic obstructive pulmonary disorder (COPD), chronic wounds and many more.
- Potential drug screening platform: High-throughput format combined with patient samples allows for screening to identify personalized, novel antibiotic treatments.
An improved AST method with high physiological relevance
Antibiotic treatments for chronic infections, such those present in cystic fibrosis (CF), often lack efficacy and result in poor clinical outcomes. The antibiotic susceptibility tests (ASTs) typically used to generate the minimum inhibitory concentration (MIC) to inform antibiotic regimens lack physiological relevance due to the inability to replicate the polymicrobial environments present at infection sites.
Dr. Ryan Hunter, Ph.D., and his team at the University of Minnesota has developed an improved AST method that features a polymicrobial testing environment derived from actual patient samples. This improved AST method is superior to current methods, which utilize only a single microbe population to determine antibiotic susceptibility. Dr. Hunter and his group have successfully demonstrated proof-of-concept using sputum samples from CF patients. This method not only allows for increased physiological relevance, but also provides a modular platform to determine antibiotic susceptibility for a variety of polymicrobial infections and sample compositions to inform personalized antibiotic regimens for patients.
Phase of Development
TRL: 3Proof-of-concept has been achieved in using sputum samples from cystic fibrosis (CF) patients.
Desired Partnerships
This technology is now available for:- License
- Sponsored research
- Co-development
Please contact our office to share your business’ needs and learn more.
Researchers
- Dr. Ryan C. Hunter Associate Professor, Department of Microbiology and Immunology
-
expand_more library_books References (1)
- Jeffrey M. Flynn, Lydia C. Cameron, Talia D. Wiggen, Jordan M. Dunitz, William R. Harcombe, Ryan C. Hunter , Disruption of Cross-Feeding Inhibits Pathogen Growth in the Sputa of Patients with Cystic Fibrosis, mSphere
-
expand_more cloud_download Supporting documents (1)Product brochureCommunity-based antibiotic screening method for polymicrobial infections (2020-090).pdf